How we read a peptide’s blueprint · ~4 min read
Precision Decoding
Edman degrades from one end; mass spectrometry weighs fragments—together they turn mystery chains into known sequences.
Why sequence matters
Function follows sequence—one substitution can change binding, stability, and immune visibility.
If you don’t know the order of amino acids, you don’t really know what you have. That’s true for research peptides, compounding ingredients, and forensic identification of unknown samples.
Edman degradation — one letter at a time
Chemically clip the N-terminus, identify it, repeat.
Like peeling tape from a roll one strip at a time and labeling each strip.
Classic Edman chemistry labels and cleaves the N-terminal residue, identifies it chromatographically, then repeats. It’s elegant but slow for long chains and struggles with blocked or modified termini.
Mass spectrometry — weigh the puzzle pieces
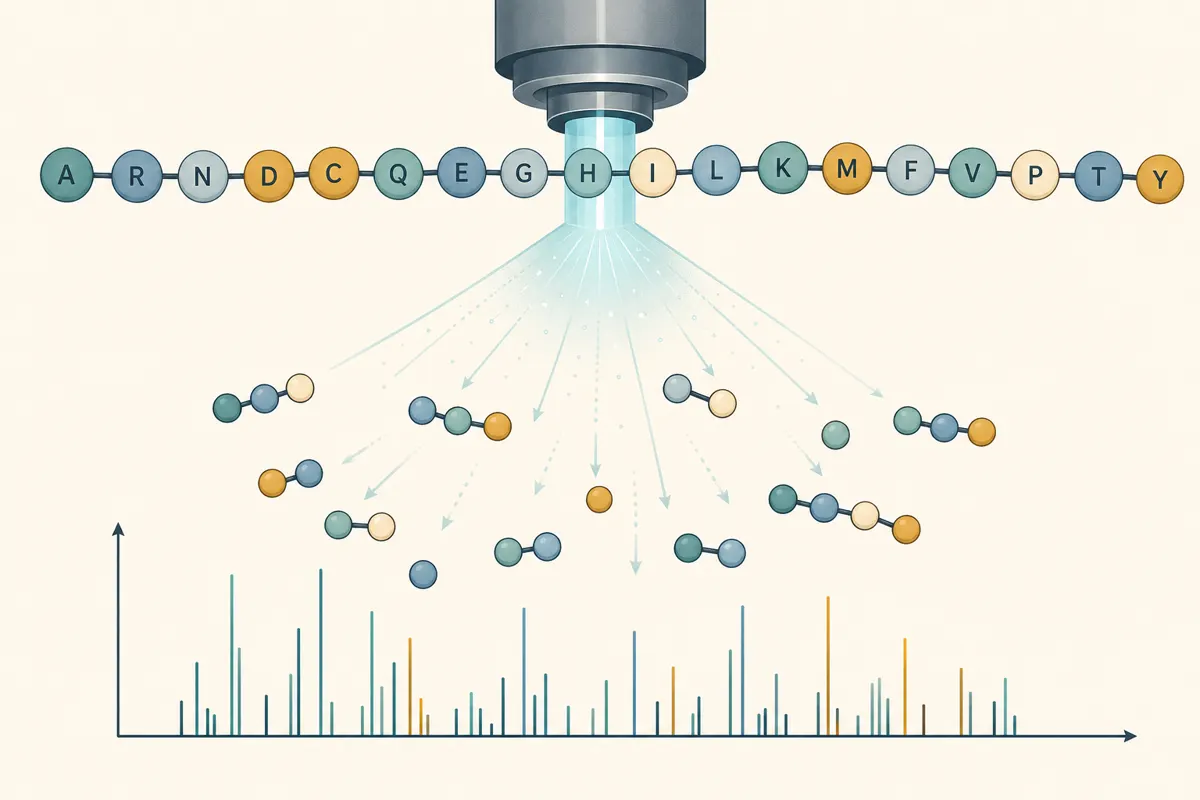
Fragment the peptide, measure fragment masses, and back-solve the sequence.
Modern workflows ionize the peptide, fragment it (often into b-ions from the N-terminal side and y-ions from the C-terminal side), and match mass-to-charge patterns to possible sequences. It’s fast and powerful but needs expertise—interpretation is where mistakes hide.